Basic R
Lesson 1 with Ian Carroll
Why learn R?
- Original design for interactive statistical analysis
- General purpose scripting for your whole pipeline
- Bleeding-edge packages expand on “base R”
- Vast community
within statistics and ecology - Open source
What is R?
- Language: a vocabulary and a syntax (with lots of punctuation!)
- Interpreter: software that evaluates expressions in the R language
The Console
The interpreter accepts commands interactively through the console.
Basic math, as you would type it on a calculator, is usually a valid command in the R language:
> 1 + 2
[1] 3
> 4^2
[1] 16
- Question
- Why is the output prefixed by
[1]? - Answer
- That’s the index, or position in a vector, of the first result.
A command giving a vector of results shows this clearly:
> seq(1, 100)
[1] 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
[18] 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34
[35] 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51
[52] 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68
[69] 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85
[86] 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100
The interpreter understands more than arithmatic operations.
That last command told it to use (or “call”) the function seq().
Most of “learning R” involves getting to know a whole lot of functions, the effect of each function’s arguments (e.g. the input values 1 and 10), and what each function returns (e.g. the output vector).
R as Calculator
A good place to begin learning R is with its built-in mathematical functionality.
Arithmatic operators
Try +, -, *, /, and ^ (for raising to a power).
> 5/3
[1] 1.666667
Logical tests
Test equality with == and inequality with =<, <, !=, >, or =>.
> 1/2 == 0.5
[1] TRUE
Math functions
Common mathematical functions like sin, log, and sqrt, exist along side some universal constants.
> sin(2 * pi)
[1] -2.449294e-16
Programming idoms
Common computer programming functions like ‘rep’, ‘sort’, and ‘range’
> rep(2, 5)
[1] 2 2 2 2 2
Parentheses
Sandwiching something with ( and ) has two possible meanings.
Group sub-expressions by parentheses on an as-needed basis.
> (1 + 2) / 3
[1] 1
Call functions by typing their name and comma-separated arguments between parentheses.
> logb(2, 2)
[1] 1
Assignment
When you start a new session, the R interpreter already recognizes many things, including
- any number
- any string of characters
- nearly universal operators (e.g.
+and/) - operators specific to R (e.g.
$and%*%) - functions in “base R”
To reference a number or function just type it in as above. To referece a string of characters, surround them in quotation marks.
> 'ab.cd'
[1] "ab.cd"
Without quotation marks, the interpreter checks for things named ab.cd and
doesn’t find anything:
> ab.cd
Error in eval(expr, envir, enclos): object 'ab.cd' not found
- Question
- Is it better to use
'or"? - Answer
- Neither one is better. You will often encounter stylistic choices like this, so if you don’t have a personal preference try to mimic existing styles.
You can expand the vocabulary known to the R interpreter by creating a new
variable. Using the symbol <- is referred to as assignment: the output of
any command to the right of <- gets the name given on its left.
> x <- seq(0, 100)
You’ll notice that nothing prints to the console, because we assigned the output to a variable.
We can print the value of x by evaluating it without assignment.
> x
[1] 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16
[18] 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33
[35] 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50
[52] 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67
[69] 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84
[86] 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100
Assigning values to new variables (to the left of a <-) is the only time you
can reference something previously unknown to the interpreter. All other
commands must reference things already in the interpreter’s vocabulary.
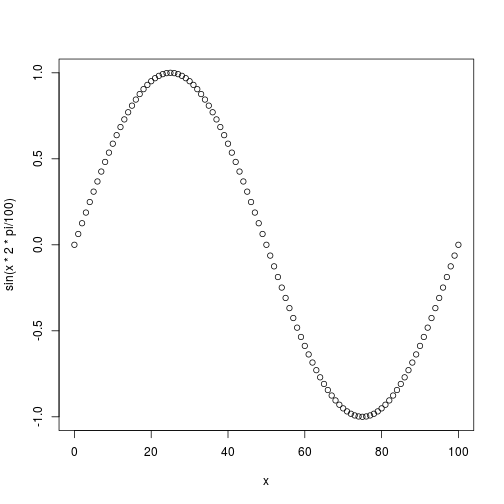
Once assigned to a variable, a value becomes known to R and you can refer to it in other commands.
> plot(x, sin(x * 2 * pi / 100))
The Environment
In the RStudio IDE, the environment tab displays the variables added to R’s vocabulary in the current session.
- Variables do not persist between sessions (unless loaded from .Rdata)
- Variables only change their value on re-assignment
The Editor
The console is for evaluating commands you don’t intend to keep or reuse. It’s useful for testing commands and poking around. The editor is where you compose scripts that will process data, perform analyses, code up visualizations, and even write reports.
These work together in RStudio, which has multiple ways to send parts of the script you are editing to the console for immediate evaluation. Alternatively you can “source” the entire script.
Open up “worksheet-1.R” in the editor, and follow along by replacing the ... placeholders with the code here. Then evalute just this line (Ctrl+Enter on Windows, ⌘+Enter on Mac OS).
vals <- seq(1, 100)
Our call to the function seq could have been much more explicit. We could give
the arguments by the names that seq is expecting.
vals <- seq(from = 1,
to = 100)
Run that code by moving your cursor anywhere within those two lines and clicking “Run”, or by using the keyboard shortcut Ctrl-Return or ⌘ Return.
- Question
- What’s an advantage of naming arguments?
- Answer
- One advantage is that you can put them in any order. A related advantage is that you can then skip some arguments, which is fine to do if each skipped argument has a default value. A third advantage is code readability, which you should always be conscious of while writing in the editor.
Readability
Code readability in the editor cuts both ways: sometimes verbosity is useful, sometimes it is cumbersome.
The seq() function has an alternative form available when only the from and
to arguments are needed.
> 1:100
[1] 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17
[18] 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34
[35] 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51
[52] 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68
[69] 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85
[86] 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100
The : operator should be used whenever possible because it replaces a common,
cumbersome function call with an brief, intuitive syntax. Likewise, the assign
function duplicates the functionallity of the <- symbol, but is never used
when the simpler operator will suffice.
Function documentation
How would you get to know these properties and the names of a function’s arguments?
> ?seq
How would you even know what function to call?
> ??sequence
Load data into R
We will use the function read.csv() that reads a Comma-Separated-Values file.
The essential argument for the function to read in data is the path to the file,
and optional additional arguments that adjust how the file is interpreted.
Additional file types can be read in using read.table(); in fact, read.csv()
is a simple wrapper for the read.table() function having set some default
values for some of the optional arguments (e.g. sep = ",").
Type read.csv( into the console and then press tab to see what arguments
this function takes. Hovering over each item in the list will show a description
of that argument from the help documentation about that function. Specify the
values to use for an argument using the syntax name = value.
> read.csv(file = "data/plots.csv", header = TRUE)
id treatment
1 1 Spectab exclosure
2 2 Control
3 3 Long-term Krat Exclosure
4 4 Control
5 5 Rodent Exclosure
6 6 Short-term Krat Exclosure
7 7 Rodent Exclosure
8 8 Control
9 9 Spectab exclosure
10 10 Rodent Exclosure
11 11 Control
12 12 Control
13 13 Short-term Krat Exclosure
14 14 Control
15 15 Long-term Krat Exclosure
16 16 Rodent Exclosure
17 17 Control
18 18 Short-term Krat Exclosure
19 19 Long-term Krat Exclosure
20 20 Short-term Krat Exclosure
21 21 Long-term Krat Exclosure
22 22 Control
23 23 Rodent Exclosure
24 24 Rodent Exclosure
- Question
- Is the
headerargument necessary? - Answer
- No. Look at
?read.csvto see thatTRUEis the default value for this argument.
Use the assignment operator “<-“ to store that data in memory and work with it
plots <- read.csv(file = "data/plots.csv")
animals <- read.csv(file = "data/animals.csv")
You can specify what indicates missing data in the read.csv function using
either na.strings = "NA" or na = "NA". You can also specify multiple things
to be interpreted as missing values, such as na.strings = c("missing", "no
data", "< 0.05 mg/L", "XX").
After reading in the “animals.csv” and “plots.csv” files, let’s explore what types of data are in each column and what kind of structure your data has.
> str(plots)
'data.frame': 24 obs. of 2 variables:
$ id : int 1 2 3 4 5 6 7 8 9 10 ...
$ treatment: Factor w/ 5 levels "Control","Long-term Krat Exclosure",..: 5 1 2 1 3 4 3 1 5 3 ...
> summary(plots)
id treatment
Min. : 1.00 Control :8
1st Qu.: 6.75 Long-term Krat Exclosure :4
Median :12.50 Rodent Exclosure :6
Mean :12.50 Short-term Krat Exclosure:4
3rd Qu.:18.25 Spectab exclosure :2
Max. :24.00
Data types
| Type | Example |
|---|---|
| double | 3.1, -4, Inf, NaN |
| integer | -4L, 0L, 999L |
| character | ‘a’, ‘4’, ‘👏’ |
| logical | TRUE, FALSE |
| missing | NA |
Data structures
Compound objects, built from one or more of these data types, or even other objects.
Common one-dimensional, array data structures:
- Vectors
- Lists
- Factors
Vectors
Vectors are the basic data structure in R. They are a collection of data that are all of the same type. Create a vector by combining elements together using the function c().
counts <- c(4, 3, 7, 5, 2)
All elements of an vector must be the same type, so when you attempt to combine different types they will be coerced to the most flexible type.
> c(1, 2, "c")
[1] "1" "2" "c"
Lists
Lists are like vectors but their elements can be of any data type or structure.
Construct lists with list() instead of c():
> list(1, 2, "c")
[[1]]
[1] 1
[[2]]
[1] 2
[[3]]
[1] "c"
Lists can even include another list!
> list(1, list(2, 3))
[[1]]
[1] 1
[[2]]
[[2]][[1]]
[1] 2
[[2]][[2]]
[1] 3
Factors
A factor is a vector that can contain only predefined values, and is used to
store categorical data. Factors are like integer vectors, but posess a levels
attribute that assigns names to however many discrete categories are specified.
Use factor() to create a vector with predefined values, which are often
characters or “strings”.
education <- factor(
c("college", "highschool", "college", "middle", "middle"),
levels = c("middle", "highschool", "college"))
The str function notes the labels, but prints the integers assigned in their
stead.
> str(education)
Factor w/ 3 levels "middle","highschool",..: 3 2 3 1 1
Tables, Matrices & Arrays
Data can be stored in several additional data structures depending on its complexity.
| Dimensions | Homogeneous | Heterogeneous |
|---|---|---|
| 1d | c() | list() |
| 2d | matrix() | data.frame() |
| nd | array() |
Of these, the data frame is far and away the most used.
Data frames
Data frames are 2-dimensional and can contain heterogenous data like numbers in one column and a factor in another.
It is the data structure most similar to a spreadsheet, with two key differences:
- The columns are equal-length vectors.
- As vectors, the columns are homogeneous and cannot hold values of the “wrong” type.
Creating a data frame from scratch can be done by combining vectors with the data.frame() function.
df <- data.frame(education, counts)
There are several functions to get to know a data frame:
dim() |
dimensions |
nrow(), ncol() |
number of rows, columns |
names() |
(column) names |
str() |
structure |
summary() |
summary info |
head() |
shows beginning rows |
> names(df)
[1] "education" "counts"
Parts of an Object
Parts of a data structure are always accessible, either by their name or by their position, using square brackets: [ and ].
Position
> counts[1]
[1] 4
> counts[3]
[1] 7
Names
Parts of an object may also have a name. The names can be given when you are creating a vector or afterwards using the names() function.
> df['education']
education
1 college
2 highschool
3 college
4 middle
5 middle
names(df) <- c('ed', 'ct')
> df['ed']
ed
1 college
2 highschool
3 college
4 middle
5 middle
- Question
- This use of
<-withnames(x)on the left is a little odd. What’s going on? - Answer
- We are overwriting an existing variable, but one that is accessed through the output of the function on the left rather than the global environment.
For a multi-dimensional array, separate the dimension along which a part is requested with a comma.
> df[3, 'ed']
[1] college
Levels: middle highschool college
It’s fine to mix names and indices when selecting parts of an object.
There are multiple ways to access several parts of an object together.
| Part | Result |
|---|---|
| positives | elements at given positions |
| negatives | given positions omitted |
| logicals | elements where the corresponding position is TRUE |
| nothing | all the elements |
days <- c(
"Sunday", "Monday", "Tuesday", "Wednesday",
"Thursday", "Friday", "Saturday")
weekdays <- days[2:6]
weekend <- days[c(1, 7)]
> weekdays
[1] "Monday" "Tuesday" "Wednesday" "Thursday" "Friday"
> weekend
[1] "Sunday" "Saturday"
Subsetting data frames
The $ sign is an operator that makes for quick access to a single, named part of an object.
It’s most useful when used interactively with “tab completion” on the columns of a data frame.
> df$ed
[1] college highschool college middle middle
Levels: middle highschool college
A logical test applied to a single column produces a vector of TRUE and FALSE values that’s the right length for subsetting the data.
> df[df$ed == 'college', ]
ed ct
1 college 4
3 college 7
Functions
Functions package up a batch of commands. There are several reasons to develop functions in R for data analysis:
- reuse
- readability
- modularity
- consistency
Writing functions to use multiple times within a project prevents you from duplicating code, a real time-saver when you want to update what the function does. If you see blocks of similar lines of code through your project, those are usually candidates for being moved into functions.
Anatomy of a function
Like all programming languages, R has keywords that are reserved for import
activities, like creating functions. Keywords are usually very intuitive, the
one we need is function.
function(...) {
...
return(...)
}
Three components:
- arguments: control how you can call the function
- body: the code inside the function
- return value: controls what output the function gives
We’ll make a function to extract the first row of its argument, which we give a name to use inside the function:
function(z) {
result <- z[1, ]
return(result)
}
Note that z doesn’t exist until we call the function, which merely contains
the instructions for how any z will be handled.
Finally, we need to give the function a name so we can use it like we used c()
and seq() above.
first <- function(z) {
result <- z[1, ]
return(result)
}
> first(df)
ed ct
1 college 4
- Question
- Can you explain the result of entering
first(counts)into the console? - Answer
- The function caused an error, which prompted the interpreter to print a helpful error message. Never ignore an error message. (It’s okay to ignore a “warning”.)
Flow control
The R interpreter’s “focus” flows through a script (or any section of code you run) line by line. Without additional instruction, every line is processed from the top to bottom. “Flow control” is the generic term for causing the interpreter to repeat or skip certain lines, using concepts like “for loops” and “if/else conditionals”.
Flow control happens within blocks of code isolated between curly braces { and }, known as “statements”.
if (...) {
...
} else {
...
}
The keyword if must be followed by a logical test which determines, at runtime, what to do next.
The R interpreter goes to the first statement if the logical value is TRUE and to the second statement if it’s FALSE.
An if/else conditional would allow the first function to avoid the error thrown by calling first(counts).
first <- function(dat) {
if (is.vector(dat)) {
result <- dat[1]
} else {
result <- dat[1, ]
}
return(result)
}
> first(df)
ed ct
1 college 4
> first(counts)
[1] 4
Review
In this introduction to R, we touched on several key parts of scripting for data analysis.
- RStudio panes
- Variable assignment
- Data structures
- Subsetting data
- Functions
- Flow control
Special characters in R
Perhaps more than most languages, an R script can appear like a jumble of archaic symbols. Here is a little table of characters to recognize as having special meaning.
| Symbol | Meaning |
|---|---|
? |
get help |
# |
comment |
: |
sequence |
::, ::: |
access namespaces (advanced) |
<- |
assignment |
$, [ ], [[ ]] |
subsetting |
% % |
infix operators, e.g. %*% |
{ } |
statements |
. |
|
@ |
slot (advanced) |
The . in R has no fixed meaning and is often used as _ might be used to
separate words in a variable name.
Exercises
Exercise 1
Use the quadratic formula to find that satisfies the equation .
Exercise 2
By default, all character data is read in to a data.frame as factors. Use the
read.csv() argument stringsAsFactors to suppress this behavior, then
subsequently modify the sex column in animals to make it a factor. Remember
that columns of a data.frame are identified to the R interpreter with the $
operator, e.g. animals$sex.
Exercise 3
Use the typeof function to inspect the data type of counts, and do the same
for another variable to which you assign a list of numbers. Why are they
different? Use c to combine counts with the new variable you just created
and inspect the result with typeof. Does c always create vectors?
Exercise 4
Create a data frame with two columns, one called “species” with four strings and
another called “abund” with four numbers. Store your data frame as a variable
called data.
Exercise 5
- Get weekdays using negative integers.
- Get M-W-F using a vector of postitions generated by
seq()that uses thebyargument (don’t forget to?seqfor help).
Exercise 6
The keywords else and if can be combined to allow flow control among more
than two statements, as below. Expand the first function once again to
differentiate between dat provided as a matrix and as a data.frame. It’s
up to you what the “first” element of a matrix should be!
if (...) {
...
} else if {
...
} else {
...
}
Solutions
Solution 1
> (-0.3 + sqrt(0.3 ^ 2 - 4 * 1.5 * -2.9)) / (2 * 1.5)
[1] 1.294035
Solution 2
animals <- read.csv('data/animals.csv', stringsAsFactors = FALSE, na.strings = '')
animals$sex <- factor(animals$sex)
> str(animals)
'data.frame': 35549 obs. of 9 variables:
$ id : int 2 3 4 5 6 7 8 9 10 11 ...
$ month : int 7 7 7 7 7 7 7 7 7 7 ...
$ day : int 16 16 16 16 16 16 16 16 16 16 ...
$ year : int 1977 1977 1977 1977 1977 1977 1977 1977 1977 1977 ...
$ plot_id : int 3 2 7 3 1 2 1 1 6 5 ...
$ species_id : chr "NL" "DM" "DM" "DM" ...
$ sex : Factor w/ 2 levels "F","M": 2 1 2 2 2 1 2 1 1 1 ...
$ hindfoot_length: int 33 37 36 35 14 NA 37 34 20 53 ...
$ weight : int NA NA NA NA NA NA NA NA NA NA ...
Solution 3
x <- list(3, 4, 5, 7)
> typeof(counts)
[1] "double"
> typeof(x)
[1] "list"
> typeof(c(counts, x))
[1] "list"
The variable x has a data type of list, so R does not restrict its elements
to a particular type as it does for vectors. The result of combining a list and
vector is a list, because the list is the more flexible data structure.
Solution 4
species <- c('ape', 'bat', 'cat', 'dog')
abund <- 1:4
data <- data.frame(species, abund)
> str(data)
'data.frame': 4 obs. of 2 variables:
$ species: Factor w/ 4 levels "ape","bat","cat",..: 1 2 3 4
$ abund : int 1 2 3 4
Solution 5
sol1 <- days[c(-1, -7)]
sol2 <- days[seq(2, 7, 2)]
> sol1
[1] "Monday" "Tuesday" "Wednesday" "Thursday" "Friday"
> sol2
[1] "Monday" "Wednesday" "Friday"
Solution 6
first <- function(dat) {
if (is.vector(dat)) {
result <- dat[1]
} else if (is.matrix(dat)) {
result <- dat[1, 1]
} else {
result <- dat[1, ]
}
return(result)
}
> m <- matrix(1:9, nrow = 3, ncol = 3)
> first(m)
[1] 1
If you need to catch-up before a section of code will work, just squish it's 🍅 to copy code above it into your clipboard. Then paste into your interpreter's console, run, and you'll be ready to start in on that section. Code copied by both 🍅 and 📋 will also appear below, where you can edit first, and then copy, paste, and run again.
# Nothing here yet!